How are FOXN1 deficiencies characterized?
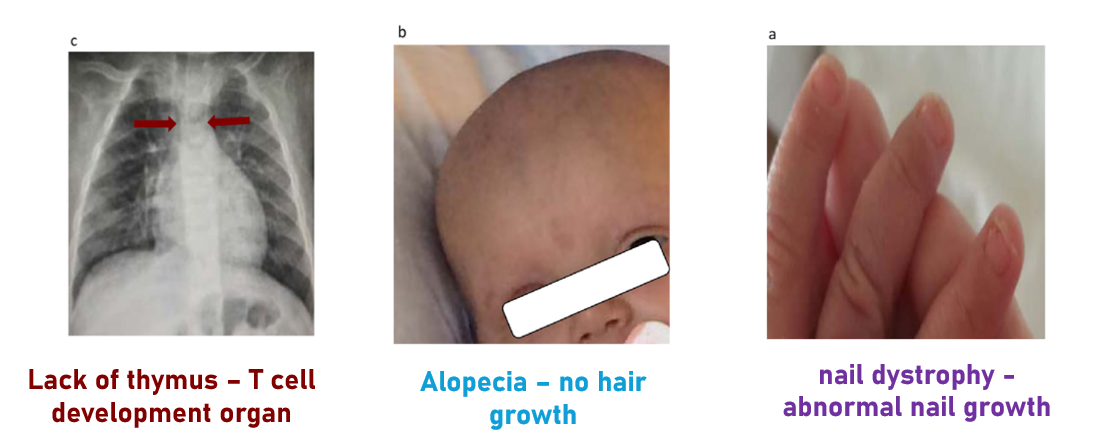
FOXN1 deficiencies are typically recognized by three main phenotypes: a lack of the thymus, alopecia, and nail dystrophy, in a disease called T cell immunodeficiency, congenital alopecia, and nail dystrophy.
Loss of FOXN1 leads to epithelial differentiation defects
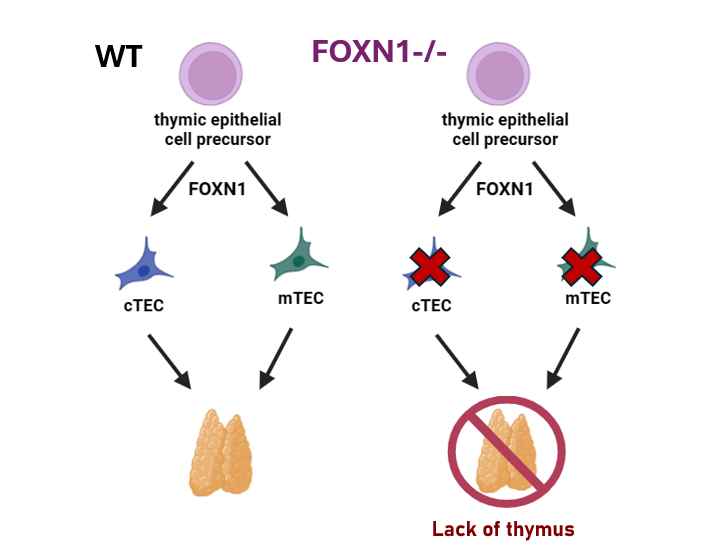
While each of the phenotypes may seem unrelated, they actually are all due to lack of epithelial cell differentiation. Mutations in the FOXN1 gene prevent important epithelial cell precursors from differentiating into epithelial cells in a variety of organs, including the example given of the thymus. Normally, FOXN1 drives thymic epithelial cell precursors to become medullary thymic epithelial cells and cortical thymic epithelial cells which together help form the thymus. When FOXN1 is mutated, thymic cell precursors do not differentiate, and the thymus does not form.
Epithelial differentiation is important to wound healing
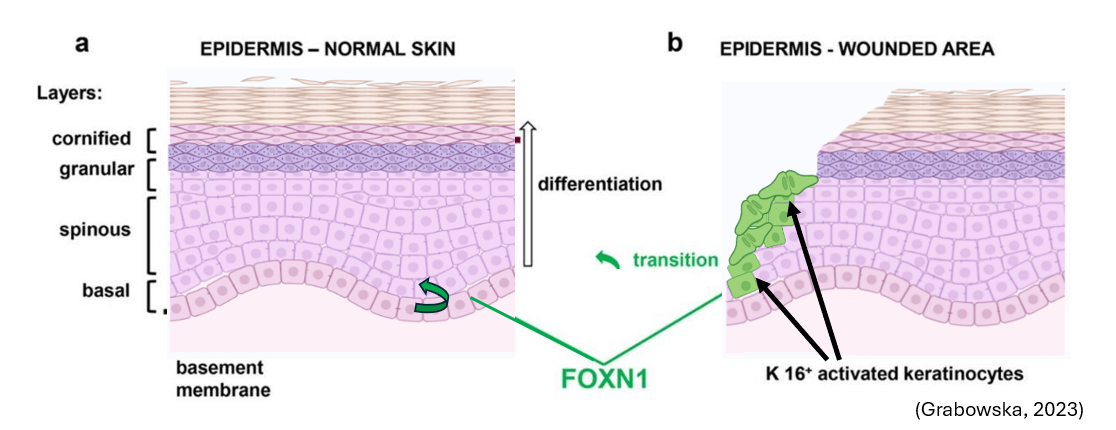
Epithelial differentiation also plays an important role in the skin. During development, FOXN1 is expressed in basal epithelial cells to create the different layers of the skin. During wound healing, keratinocytes on the wound edge also express FOXN1 is also expressed. One interesting phenotype, however, is that rather than wound healing being disrupted by mutations in FOXN1, it happens at a faster rate.
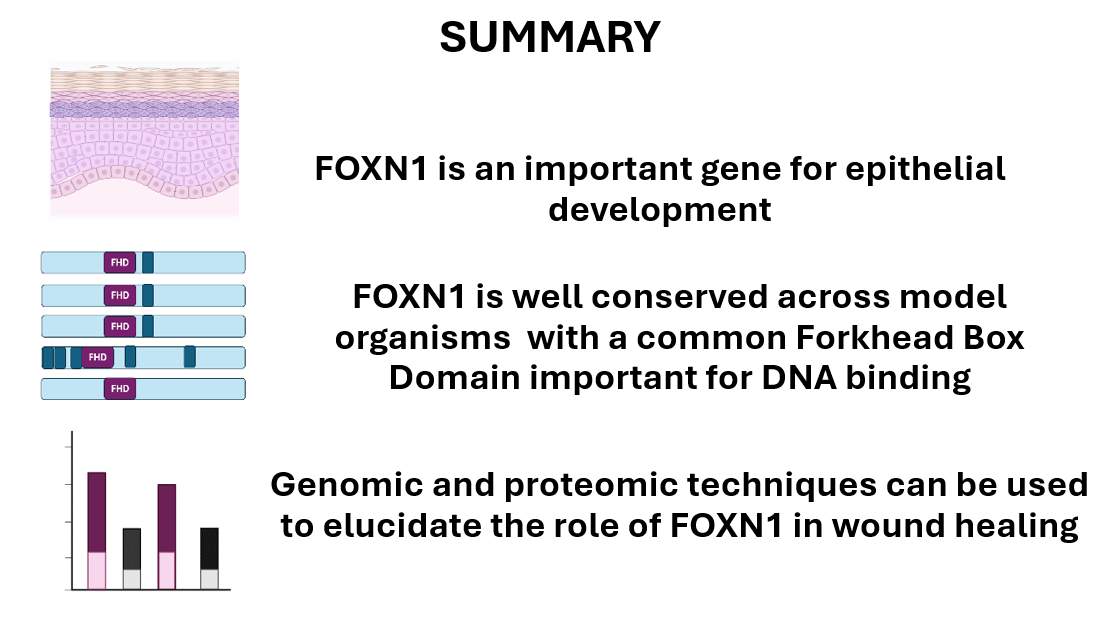
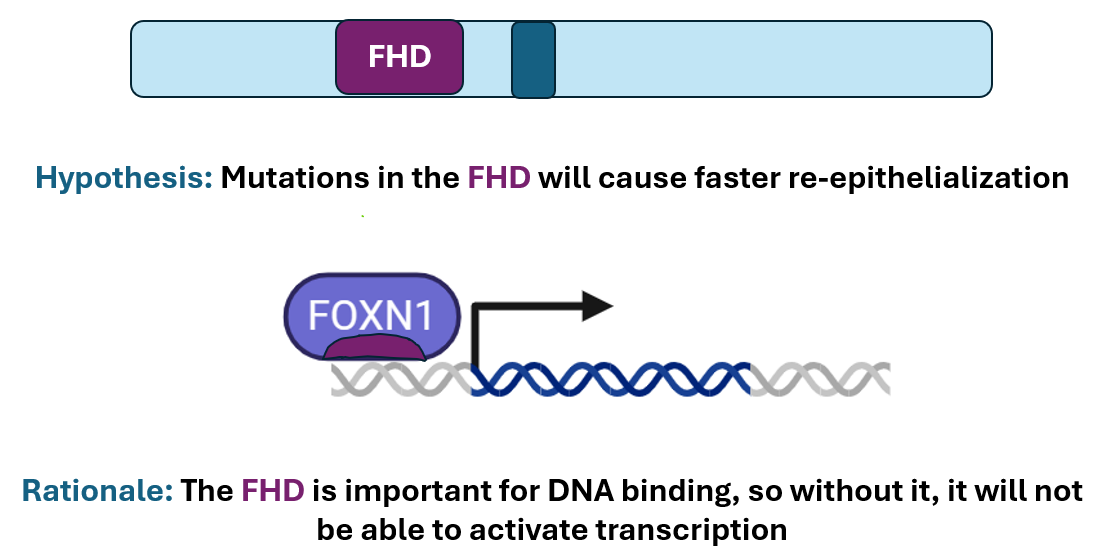
FOXN1’s forkhead domain drives epithelial cell differentiation
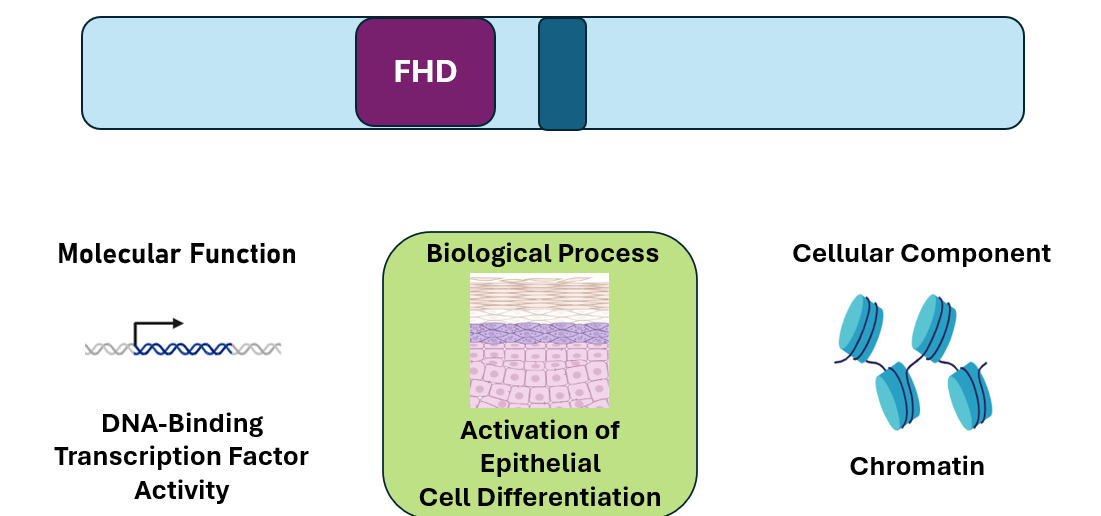
The Forkhead Domain is important for transcription factor activities. This is supported by the GO terms found using Ami GO. The GO terms DNA-binding transcription factor activity, activation of epithelial cell differentiation and chromatin all support that FOXN1 plays a role in regulation of epithelial cell differentiation through gene regulation.
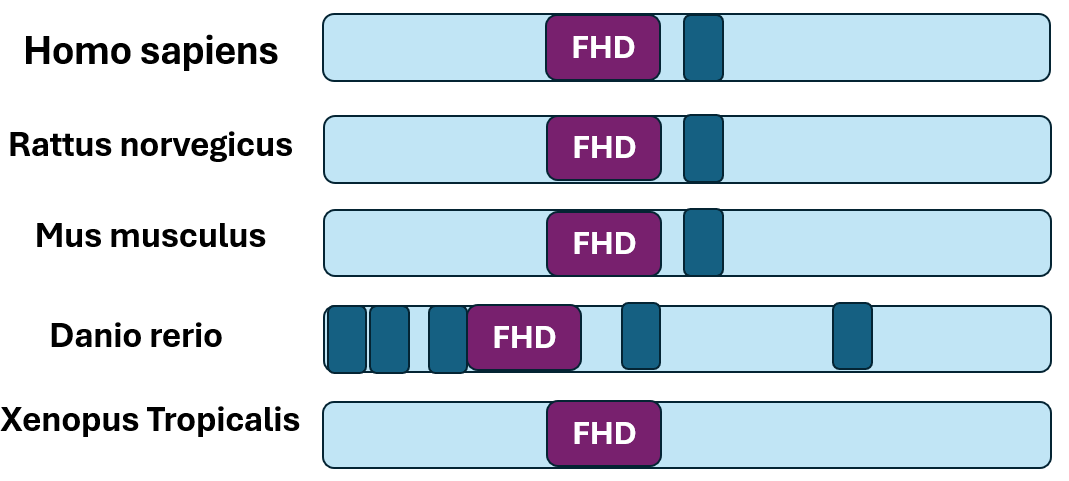
The Forkhead Domain is conserved across species
Domain analysis of model organism FOXN1 orthologs using SMART revealed conservation of the Forkhead Box Domain. This implies that this domain is important to the function of the gene across species.
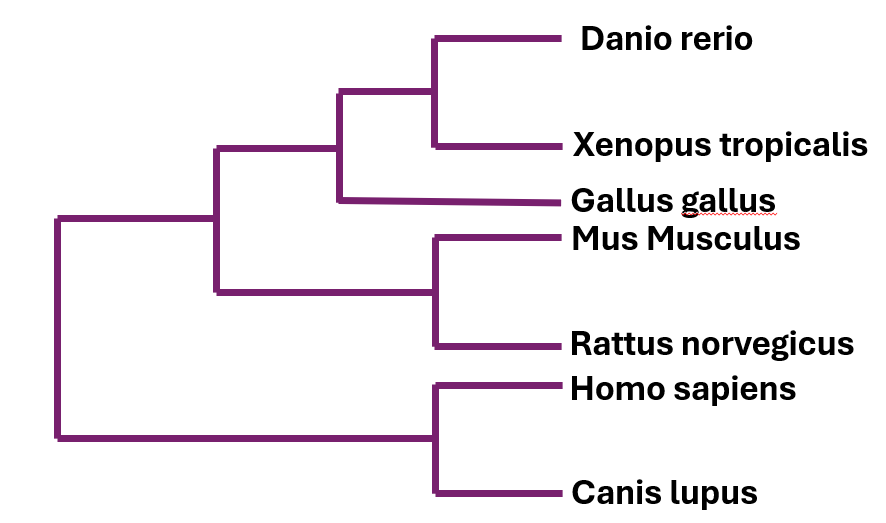
FOXN1 is only found in vertebrates
The phylogenetic tree of the FOXN1 tree shows that mammals have a more similar protein to one another, however because this protein is shared among vertebrates with high conservation, this is not a reason to rule out other vertebrate model organisms.
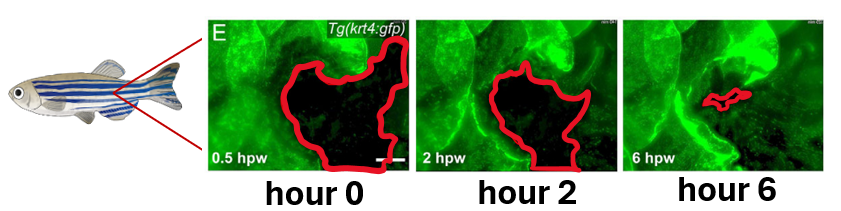
Zebrafish is a good model organism for wound healing and development
Zebrafish are a good model to study wound healing because they can easily be assayed using transgenic fish, which make it easier to distinguish between cells on the wound edge. Wound healing assays work by measuring the post-wound area over time. Additionally, zebrafishes' natural accelerated healing allows for these assays to be completed quicker.
Because they can be monitored throughout their entire lives, zebrafish are great models for development.
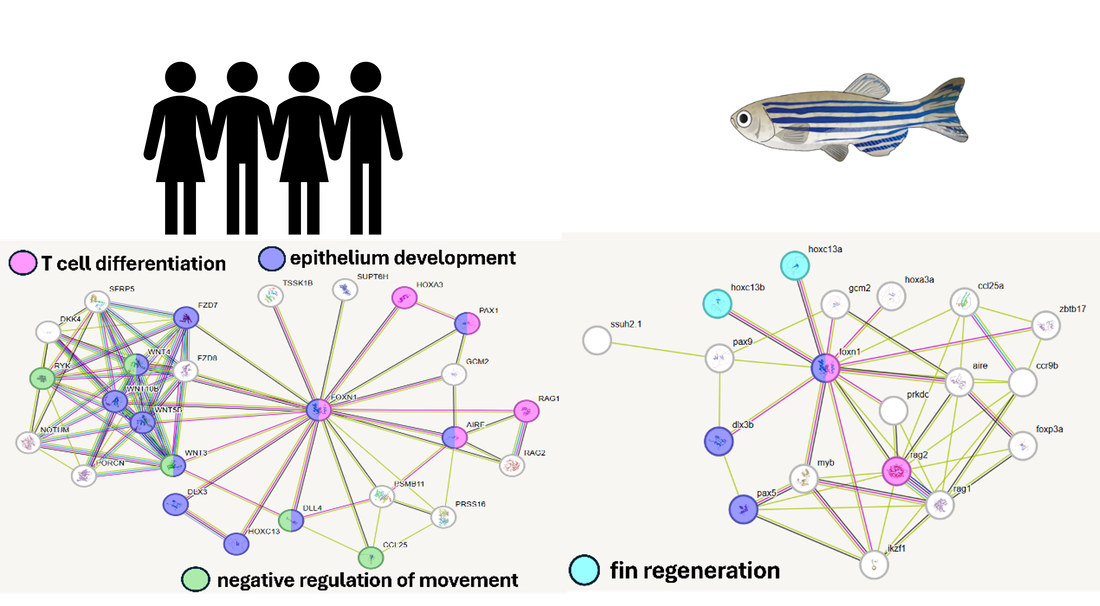
Human and zebrafish FOXN1 have similar interaction networks
Zebrafish and human FOXN1 share many protein interactions. These interactions are with proteins involved in epithelium development and skin regeneration in zebrafish in addition to T cell differentiation.
GAP: How does FOXN1 regulate epithelial cell differentiation in skin?
Goal: elucidate mechanisms related to epithelial differentiation involved in FOXN1 deficiency rapid wound healing
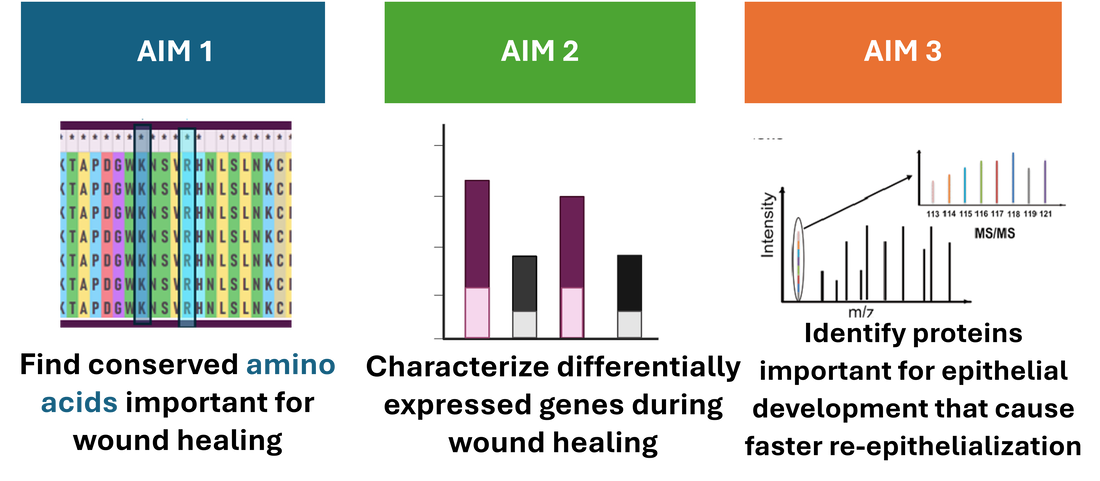
Aim 1: Find conserved amino acids important for wound healing
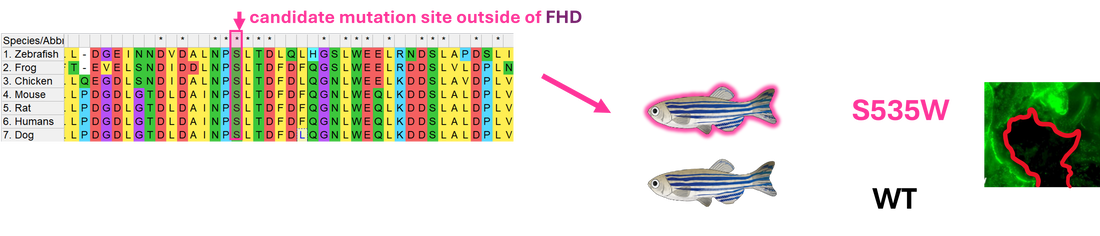
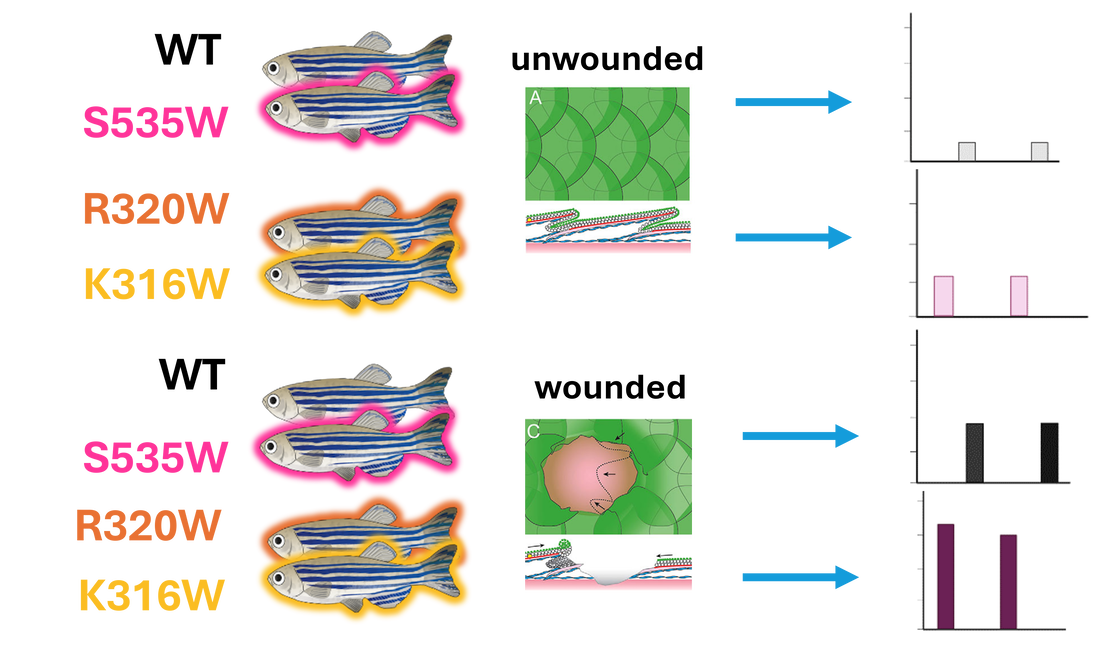
I will use NCBI BLAST to identify orthologs of FOXN1 within other organisms and use Clustal Omega to align their sequences to determine conserved amino acids across species within the Forkhead Domain and the overall protein. CRISPR-Cas9 will then be used to create mutations in the known mutation site R320W, another in the FHD, K316W, and one outside the FHD, S535W. To validate the model, a wound healing assay will be completed on each mutant and WT, determining the change in wound area throughout time.
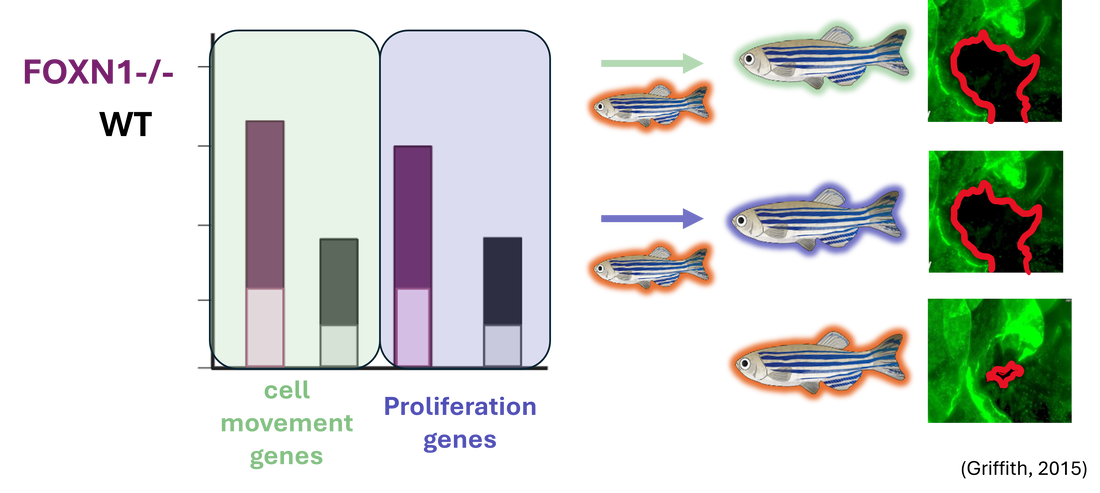
AIM 2: Identify differentially expressed genes in FOXN1 mutants during wound healing using RNA-seq
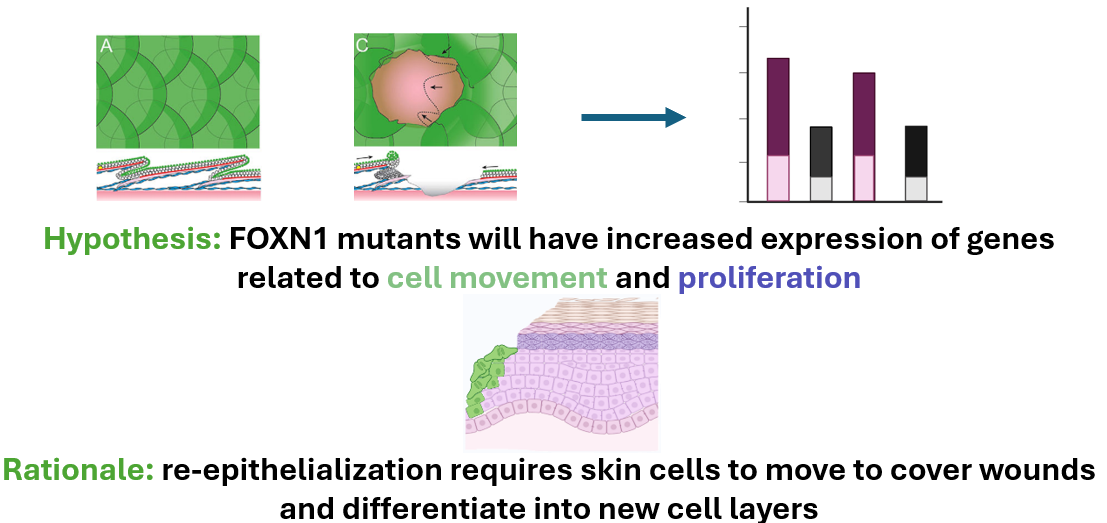
This approach will use validated mutant and control zebrafish. We will take epidermal samples from unwounded zebrafish and wounded (at the wound edge) zebrafish and will utilize RNA-sequencing to determine differential gene expression.
Relative gene expression from the validated FOXN1-deficient zebrafish will be compared to that of the WT. Differentially expressed genes will then be sorted by GO. Differentially expressed genes found to be involved in cell movement and proliferation will then be mutated in FOXN1-deficient zebrafish using CRISPR-Cas9 and a wound healing assay will be performed to validate these genes’ roles in re-epithelialization.
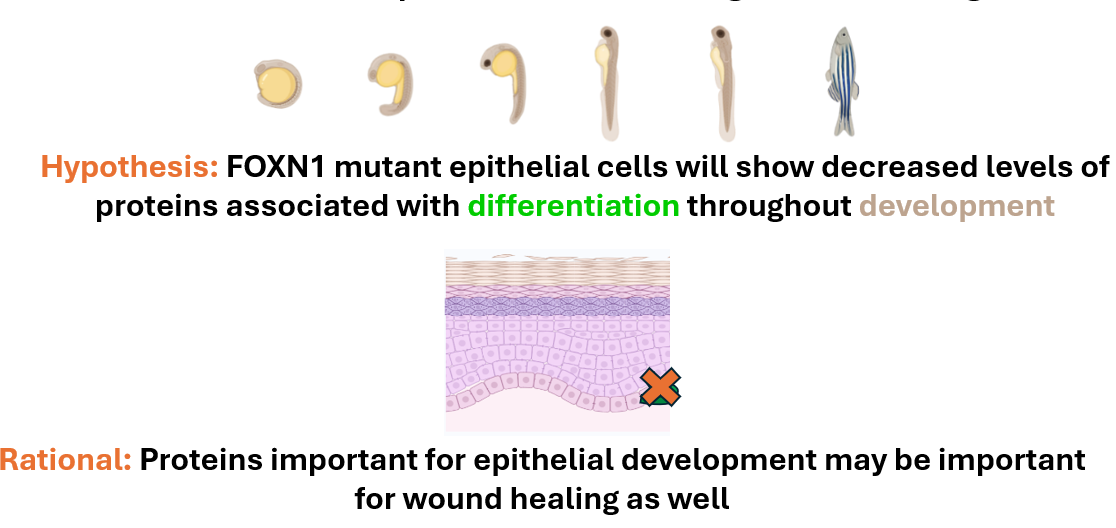
AIM 3: Identify proteins important for epithelial development that cause faster re-epithelialization during wound healing
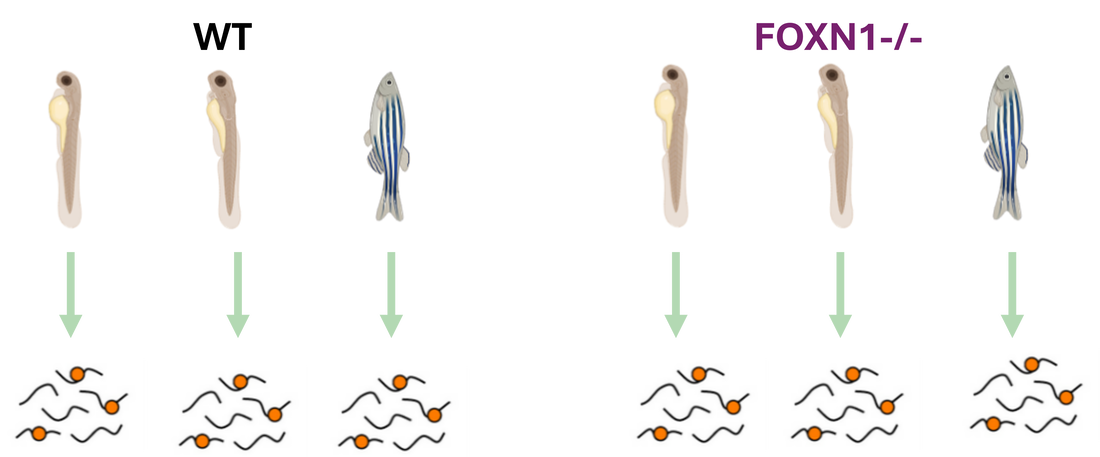
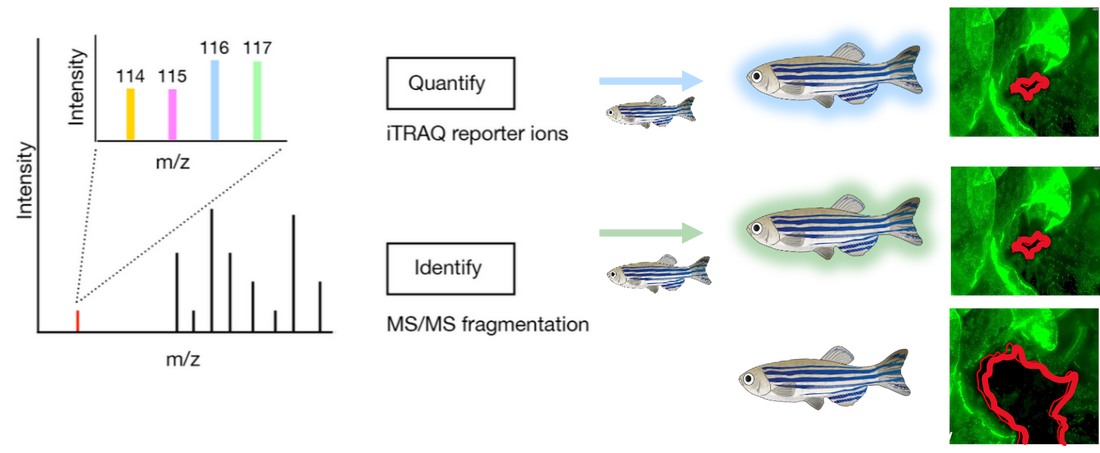
This approach will also use the validated mutant and control zebrafish lines. Epithelial samples will be taken from each zebrafish line at 2dpf, 6dpf, and 12dpf. These cells will then be independently digested and labeled using iTRAQ.
They will then be analyzed using mass-spectrometry to determine the proteins present and sorted by GO. Differentially expressed proteins found to be involved in epithelial cell differentiation will then be mutated using CRISPR-Cas9 and validated using the wound healing assay described previously.
Presentation Rough Drafts:
| zwick2292024Draft1.ppt |
| zwick3212024draft2.pptx |
| foxn1_presentation.pptx |
| foxn1_final_presentation.pdf |
Sources
PDF) A Novel FOXN1 Variant Is Identified in Two Siblings with Nude Severe Combined Immunodeficiency (researchgate.net)
Genetic interplays between Msx2 and Foxn1 are required for Notch1 expression and hair shaft differentiation – ScienceDirect
Regulatory T cells in skin regeneration and wound healing | Military Medical Research | Full Text (biomedcentral.com)
Foxn1 in Skin Development, Homeostasis and Wound Healing - PMC (nih.gov)
Impairment of the Hif-1α regulatory pathway in Foxn1-deficient (Foxn1-/- ) mice affects the skin wound healing process - PubMed (nih.gov)
Adult Zebrafish as a Model System for Cutaneous Wound-Healing Research – ScienceDirect
FOXN1 Transcription Factor in Epithelial Growth and Wound Healing: Molecular and Cellular Biology: Vol 37, No 17 (tandfonline.com)
content (mpg.de)
RNA-Seq Expression Analysis: 5 Dirty Secrets - Genevia Technologies
Informatics for RNA Sequencing: A Web Resource for Analysis on the Cloud | PLOS Computational Biology
Biorender
Genetic interplays between Msx2 and Foxn1 are required for Notch1 expression and hair shaft differentiation – ScienceDirect
Regulatory T cells in skin regeneration and wound healing | Military Medical Research | Full Text (biomedcentral.com)
Foxn1 in Skin Development, Homeostasis and Wound Healing - PMC (nih.gov)
Impairment of the Hif-1α regulatory pathway in Foxn1-deficient (Foxn1-/- ) mice affects the skin wound healing process - PubMed (nih.gov)
Adult Zebrafish as a Model System for Cutaneous Wound-Healing Research – ScienceDirect
FOXN1 Transcription Factor in Epithelial Growth and Wound Healing: Molecular and Cellular Biology: Vol 37, No 17 (tandfonline.com)
content (mpg.de)
RNA-Seq Expression Analysis: 5 Dirty Secrets - Genevia Technologies
Informatics for RNA Sequencing: A Web Resource for Analysis on the Cloud | PLOS Computational Biology
Biorender