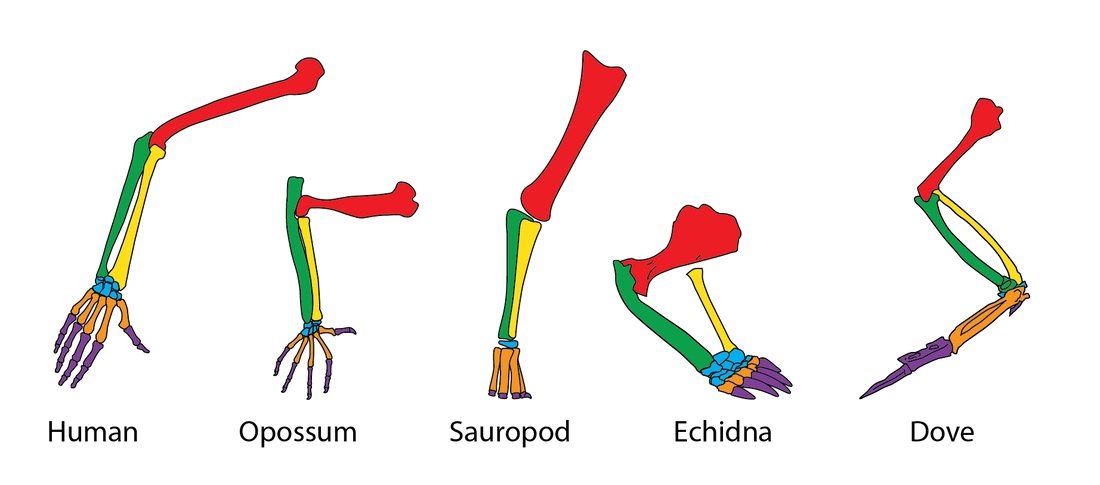
What is Homology?
Homology is the presence of similar biological structures in different species descended from a common ancestor [1]. While bones as seen above may be homologous to one another. This homology happens due to the presence of genes passed down from the common ancestor called homologs.
There are two kinds of homologs: paralogs and orthologs. Paralogs are genes that are identical as a result of a duplication event while orthologs are identical genes across species that come about directly from the common ancestor.
Orthologs between species with known protein sequences can be found through the use of NCBI BLAST, which finds regions of similarity between sequences, comparing them to databases, and calculating their statistical significance [2]. Orthologs between humans and model organisms are especially of interest as it can be informative of potential models for changes in these genes.
Human FOXN1 protein
|
The human FOXN1 gene was ran through BLASTp to find homologs in model organisms. While the gene was only found in vertebrate models, other organisms have the paralog, FOXN4.
|
Model Organisms
Discussion
The dog, zebrafish, rat, chicken, mouse and frog model organisms were found to have orthologs of the FOXN1 gene. This would make them all plausible candidates for genetic research on FOXN1.
Sources
1. National Institutes of Health. (n.d.). Homology: Orthologs and paralogs. U.S. National Library of Medicine. https://www.nlm.nih.gov/ncbi/workshops/2023-08_BLAST_evol/ortho_para.html
2. U.S. National Library of Medicine. (n.d.). Blast: Basic local alignment search tool. National Center for Biotechnology Information. https://blast.ncbi.nlm.nih.gov/Blast.cgi
IMAGES:
Biorender
This web page was produced as an assignment for Genetics 564, a capstone course at UW-Madison.